Sandeep Kumar S1#, Shalini N Swamy1#, Premalatha C S2, Pallavi V R3, Ramesh Gawari1*
1Department of Biochemistry, Kidwai Memorial Institute of Oncology, Bangalore, India
2Department of Pathology, Kidwai Memorial Institute of Oncology, Bangalore, India
3Department of Gyneconcology, Kidwai Memorial Institute of Oncology, Bangalore, India
*Corresponding author: Ramesh Gawari, Department of Biochemistry, Kidwai Memorial Institute of Oncology, Bangalore, India
# - The authors contributed equally to the work
Received: 15 July 2020; Accepted: 22 July 2020; Published: 27 July 2020
Ovarian cancer is the third leading gynecological malignancy in India with a high mortality rate. The high mortality of the disease is attributed to the late stage diagnosis of the disease. Silencing or down regulation of Tumor Suppressor genes is known to be an early event during tumorigenesis and is known to regulate several cellular events including DNA damage, proliferation and apoptosis. In this study, we have evaluated the mRNA expression of three TSG-RASSF1a, APC and GSTP1 in ovarian carcinoma tissues. We report here that 72/76 samples (94.73%) samples show a down regulation of RASSF1a gene, 47/76 (61.84%) sample show down regulation of APC and 51/76 (67.10%) samples showed down regulation of GSTP1. 100% of the benign samples showed downregulation of RASSF1a and GSTP1 and 77.77% of benign samples showed downregulation of APC gene, which indicates that the down regulation of these TSGs happen early in tumorigenesis and contribute to disease progression.
RASSF1a; APC; GSTP1; Epithelial Ovarian Cancer; Diagnosis; Gene expression
RASSF1a articles, APC articles, GSTP1 articles, Epithelial Ovarian Cancer articles, Diagnosis articles, Gene expression articles
Introduction
Ovarian Carcinoma (OC) arising from the ovarian surface epithelium accounts to about 95% of all ovarian malignancies. It is predicted that annually worldwide, 230 000 women will be diagnosed and 150 000 will die of the disease implicating a high death to incidence ratio. It stands as the seventh most commonly diagnosed cancer among women in the world and the third leading gynecological malignancy in India, with 46% survival rate 5 years after the diagnosis [1-3]. The main factor for increased mortality for EOC is the late stage detection of the disease. The survival rate in patients with early stage presentation is 92% in contrast to 29% survival rate in patients with late stage presentation [3,4]. Unfortunately over 75% of the patients are diagnosed at late stages where the disease would have already metastasized within the peritoneal cavity. Diagnosis of ovarian cancer includes the estimation of serum CA125 and pelvic ultrasonography. There are no molecular methods available for the precise detection of the disease in the early stages [1].
Inactivation or silencing of tumor suppressor genes (TSG) is known to be an early event in the initiation and progression of many cancers including EOC. TSGs are known to be involved in coding growth constraining proteins whose loss of function leads to deregulated cellular proliferation leading to tumorigenesis [5,6]. It has been reported that there are about 1217 human TSGs that are identified whose silencing is known to contribute to tumorigenesis7. Along with their growth constraining function, Tumor suppressor genes are known to be involved in several cellular processes such as DNA damage repair, induction of apoptosis and suppression of metastasis.
RASSF1a is a TSG located on chromosome 3p21.3, encodes for the Ras-super family of GTP-ases that are essential for intracellular signal transduction. RASSF1a modulates several cellular pathways including apoptotic pathway and tubulin dynamics. RASSF1a is also known to cause the repression of cyclin A and cyclin D1 to mediate cell cycle arrest [3,8,9]. RASSF1a is known to be silenced via epigenetic mechanism rather than mutational events. The silencing of this gene is known to be attributed in several cancer including lung, breast, prostate, renal and ovarian cancers [10-16].
APC1 located on chromosome 5q21-q22 is found to be expressed in many fetal and adult epithelial cells [17]. APC1 is known to play important roles in cell migration, adhesion, cytoskeleton organization and regulation of cellular proliferation [18,19]. Reports also suggest that APC1 is also involved in apoptosis and cell cycle arrest from Go/G1 to S phase [20,21]. Genetic alterations such as mutations are known to cause silencing of APC1 in many cancer types [22,23]. However, in ovarian cancer APC1 gene silencing is attributed to epigenetic silencing through promoter hypermethylation [18,24-26].
TheGSTP1gene is localized on chromosome 11q13 is a cytosolic detoxifying enzyme playing a critical role in maintaining cell integrity and protecting DNA from genotoxic stress. GSTP1 is also attributed to regulate the apoptosis signalling pathway. It has been reported that loss of GSTP1 expression, enhances cells to acquire additional genetic alterations that contribute to cancer progression.
There stands an indispensible need to diagnose ovarian cancer at the initial stages which could there by contribute to better management of the disease in addition to identifying new therapeutic targets for treatments. In the present study, we have assessed the mRNA expression of RASSF1a, APC1 and GSTP1 in primary ovarian cancer, low malignant potential and benign tissues in an attempt to assess if these could act as potential markers for diagnosis and prognosis.
Methods
Patient Samples
76 Ovarian tumor tissue was obtained (51 malignant, 16 Low Malignant Potential-LMP and 9 benign cyst adenoma) from patients reporting to Kidwai Memorial Institute of Oncology, Bangalore, India. All the samples were verified histologically by a pathologist before including in the study. Histological classification was established as per the WHO criteria and staged according to the FIGO classification. Normal ovarian tissue was obtained from patients undergoing salphingo-oopherctomy. The study was approved by the institutional scientific review board and medical ethics committee and informed consent was obtained from all patients prior to sample collection.
RNA isolation and cDNA preparation
Tumor samples were collected in RNAlater™ after histopathological confirmation. Total RNA was extracted from 30 mg of tissue using RNeasy mini kit (Qiagen, CA, USA) following the manufacturer’s protocol. The RNA was quantified using Eppendorf biospectrophotometer kinetics™ and 1μg of RNA was used for cDNA preparation. cDNA was prepared using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, CA, USA). The conditions for cDNA preparation used are mentioned in table 1.
|
Step1 |
Step2 |
Step3 |
Step4 |
|
|
Temperature |
250C |
370C |
850C |
40C |
|
Time |
10mins |
120mins |
5mins |
∞ |
Table 1: cDNA conversion
Gene Expression Analysis
Gene expression of RASSF1a, APC1 and GSTP1 was quantified by TaqMan chemistry on StepOnePlus™ Real-Time PCR system (Applied Biosystem, CA, USA). Gene-specific primers and probe of RASSF1a (Hs00200394_m1), APC1 (Hs01568282_m1) and GSTP1 (Hs00168310_m1) were available as TaqMan gene expression assays (Applied Biosystems). The 18S rRNA (Hs99999901_s1) was amplified and was used as an endogenous control in the quantification. The gene expression was expressed as fold change. The gene expression values were correlated with various clinicopathological parameters.
Statistical Analysis
Chi square test and Fischer exact test (two- tailed) was used for analysis and a p-value <0.05 was considered to be statistically significant.
Results
Gene Expression Analysis of RASSF1a, APC and GSTP in Ovarian Cancer
Gene Expression of RASSF1a
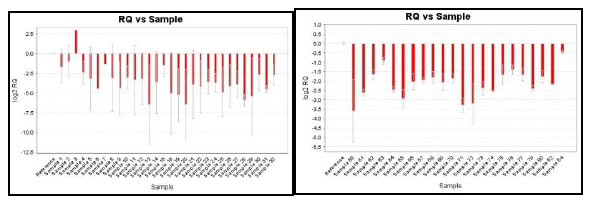
Gene expression analysis of revealed that 72 cases of the total 75 samples analyzed had down regulation of RASSF1a. RASSF1a was downregulated in 48/51 (94.10%) malignant samples, 15/16 (93.75%) LMP and 9/9 (100%) benign samples analyzed. The samples were analyzed for statistical significance using Fischer exact test and the data correlated significantly among the group compared. The above data is summarized in table2. The gene expression plot for RASSF1a of tumor samples in comparison to normal samples is depicted below.
|
Expression of RASSF1a |
Tumor type |
|||
|
Malignant (51) |
LMP (16) |
Benign(09) |
Normal (5) |
|
|
≥ 1 RQ |
03/51 (5.88%) |
1/16(6.25%) |
0/9(0%) |
5/5(100%) |
|
<1 RQ |
48/51 (94.11%) |
15/16 (93.75%) |
9/9(100%) |
0/5 |
|
p-Value |
<0.0001* |
0.0003* |
0.0005* |
|
Table 2: Representation of RASSf1a gene expression in malignant, LMP, benign and normal tissue.
Gene expression of APC
APC expression was downregulated in 47/76 samples analyzed. 31/51(60.70%) malignant, 9/16 (56.25%) LMP and 7/9 (77.77%) benign samples showed downregulation of the APC1 mRNA. The p-value was found to be 0.014 for Malignant, 0.045 for LMP, 0.021 for benign samples and is represented in table3.
The gene expression plot of APC1 gene in tumor in comparison with normal samples is shown below.
|
Expression of APC |
Tumor type |
|||
|
Malignant (51) |
LMP (16) |
Benign(09) |
Normal (5) |
|
|
≥ 1 RQ |
20/51 (39.21%) |
07/16(43.7%) |
02/9(22.22%) |
5/5(100%) |
|
<1 RQ |
31/51(60.70%) |
9/16 (56.2%) |
7/9(77.77%) |
0/5 |
|
p-Value |
0.014* |
0.045* |
0.021* |
|
Table 3: Representation of APC gene expression in malignant, LMP, benign and normal tissue
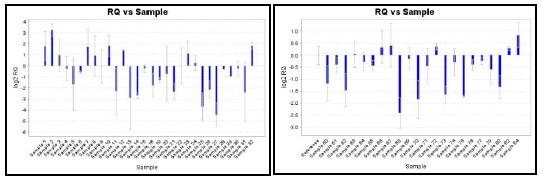
Gene Expression of GSTP1
51/76 samples showed down regulation of the GSTP1 gene, of which 29/51 (56.80%) malignant, 13/16 (81.25%) LMP and 9/9 (100%) benign samples showed down regulation. The gene expression plot of GSTP1 in tumor and normal samples is depicted in the plot below. The data with p-value is represented in Table 4.
|
Expression of GSTP |
Tumor type |
|||
|
Malignant (51) |
LMP (16) |
Benign(09) |
Normal (5) |
|
|
≥ 1 RQ |
22/51 (43.13%) |
03/16(18.75%) |
0/9(0%) |
5/5(100%) |
|
<1 RQ |
29/51(56.80%) |
13/16 (81.25%) |
9/9 (100%) |
0/5 |
|
p-Value |
0.021* |
0.002* |
<0.001* |
|
Table 4: Representation of GSTP1 gene expression in malignant, LMP, Benign and normal tissue.
37 of the 76 (48.68%) samples analyzed had low expression for all the three genes. 23 samples showed low expression of at least two genes (Table 5). This suggests that the silencing of TSG is an important contributor for tumor formation and progression.
|
SAMPLE |
RASSF1A |
APC |
GSTP |
|
1 |
LOW |
HIGH |
LOW |
|
2 |
LOW |
HIGH |
LOW |
|
3 |
HIGH |
HIGH |
HIGH |
|
4 |
LOW |
LOW |
HIGH |
|
5 |
LOW |
LOW |
HIGH |
|
6 |
LOW |
LOW |
HIGH |
|
7 |
LOW |
HIGH |
HIGH |
|
8 |
LOW |
HIGH |
HIGH |
|
9 |
LOW |
LOW |
HIGH |
|
10 |
LOW |
HIGH |
HIGH |
|
11 |
LOW |
LOW |
HIGH |
|
12 |
LOW |
HIGH |
HIGH |
|
13 |
LOW |
LOW |
LOW |
|
14 |
LOW |
LOW |
LOW |
|
15 |
LOW |
LOW |
LOW |
|
16 |
LOW |
LOW |
LOW |
|
17 |
LOW |
LOW |
LOW |
|
18 |
LOW |
LOW |
LOW |
|
19 |
LOW |
LOW |
LOW |
|
20 |
LOW |
HIGH |
LOW |
|
21 |
LOW |
HIGH |
LOW |
|
22 |
LOW |
HIGH |
LOW |
|
23 |
LOW |
LOW |
HIGH |
|
24 |
LOW |
LOW |
LOW |
|
24 |
LOW |
LOW |
LOW |
|
26 |
LOW |
LOW |
LOW |
|
27 |
LOW |
LOW |
LOW |
|
28 |
LOW |
LOW |
HIGH |
|
29 |
LOW |
LOW |
HIGH |
|
30 |
LOW |
HIGH |
HIGH |
|
31 |
LOW |
LOW |
LOW |
|
32 |
LOW |
HIGH |
HIGH |
|
33 |
LOW |
HIGH |
HIGH |
|
34 |
LOW |
HIGH |
LOW |
|
35 |
LOW |
LOW |
LOW |
|
36 |
LOW |
HIGH |
LOW |
|
37 |
LOW |
LOW |
LOW |
|
38 |
LOW |
HIGH |
HIGH |
|
39 |
LOW |
HIGH |
LOW |
|
40 |
LOW |
LOW |
LOW |
|
41 |
LOW |
LOW |
LOW |
|
42 |
LOW |
LOW |
LOW |
|
43 |
LOW |
LOW |
LOW |
|
44 |
LOW |
LOW |
LOW |
|
45 |
LOW |
LOW |
LOW |
|
46 |
LOW |
LOW |
LOW |
|
47 |
HIGH |
HIGH |
HIGH |
|
48 |
LOW |
HIGH |
HIGH |
|
49 |
LOW |
HIGH |
HIGH |
|
50 |
HIGH |
HIGH |
HIGH |
|
51 |
LOW |
LOW |
HIGH |
|
52 |
LOW |
HIGH |
HIGH |
|
53 |
LOW |
HIGH |
HIGH |
|
54 |
LOW |
HIGH |
HIGH |
|
55 |
LOW |
LOW |
LOW |
|
56 |
LOW |
LOW |
LOW |
|
57 |
LOW |
LOW |
LOW |
|
58 |
LOW |
HIGH |
LOW |
|
59 |
LOW |
LOW |
LOW |
|
60 |
LOW |
LOW |
LOW |
|
61 |
LOW |
HIGH |
LOW |
|
62 |
LOW |
HIGH |
LOW |
|
63 |
LOW |
LOW |
LOW |
|
64 |
LOW |
LOW |
LOW |
|
65 |
LOW |
LOW |
LOW |
|
66 |
LOW |
LOW |
LOW |
|
67 |
LOW |
HIGH |
LOW |
|
68 |
LOW |
LOW |
LOW |
|
69 |
LOW |
LOW |
LOW |
|
70 |
LOW |
LOW |
LOW |
|
71 |
LOW |
LOW |
LOW |
|
72 |
LOW |
LOW |
LOW |
|
73 |
LOW |
LOW |
LOW |
|
74 |
LOW |
LOW |
LOW |
|
75 |
LOW |
HIGH |
LOW |
|
76 |
LOW |
HIGH |
LOW |
Table 5: Expression profile of RASSF1a, APC1 and GSTP1 in the samples analyzed
High: ≥ 1 RQ value; Low: <1 RQ value.
Correlation of Gene Expression with Clinicopathological Parameters
The expression profiles of RASSF1a did not statistically correlate with any of the clinicopathological parameters analyzed despite the high percentage of downregulation of the gene. However, down regulation of APC1 correlated with the menopausal status with a p-value of 0.0082. GSPT1 downregulation statistically correlated with the FIGO stage of the samples analyzed (Table 6).
|
Characteristics |
N |
RASSF1A (silenced) |
APC1 (silenced) |
GSTP1 (silenced) |
|
|
Ovarian tumors |
76 |
||||
|
Menopause State |
Pre Menopause |
25 |
24/25 |
17/25 |
16/25 |
|
Post Menopause |
51 |
49/51 |
32/51 |
38/51 |
|
|
p-Value |
p = 0.986837 |
p= 0.00823608 |
p = 0.342529 |
||
|
Histological type |
Serous |
43 |
40/43 |
28/43 |
26/43 |
|
Mucinous |
02 |
2/2 |
1/2 |
½ |
|
|
Clear cell |
04 |
4/4 |
3/4 |
2/4 |
|
|
Endometriod |
02 |
2/2 |
1/2 |
½ |
|
|
p-Value |
p = 0.898028 |
p = 0.90381 |
p = 0.959089 |
||
|
FIGO Stage |
I-II |
21 |
20/21 |
11/21 |
4/21 |
|
III-IV |
30 |
28/30 |
21/30 |
21/30 |
|
|
p-Value |
p = 0.776011 |
p = 0.200259 |
p=0.000340569 |
||
|
GRADE |
I-II |
17 |
16/17 |
11/17 |
7/17 |
|
III-IV |
34 |
32/34 |
22/34 |
23/34 |
|
|
p-Value |
P=1.0 |
P=1 |
p = 0.07019 |
||
|
CA125(U/ml) |
0-35 |
14 |
14/14 |
9/14 |
11/14 |
|
35-500 |
32 |
30/32 |
19/32 |
20/32 |
|
|
500-1000 |
15 |
14/15 |
10/15 |
11/15 |
|
|
>1000 |
15 |
15/15 |
11/15 |
11/15 |
|
|
p-Value |
p = 0.586679 |
p = 0.823271 |
p = 0.68326 |
||
|
Ascites |
Presence of ascites |
57 |
55/57 |
34/57 |
40/57 |
|
Absence of ascites |
19 |
18/19 |
13/19 |
11/19 |
|
|
p-Value |
p = 0.733771 |
p = 0.495454 |
p = 0.323785 |
Table 6: Gene expression profile and correlation with clinicopathological parameters and clinicopathological parameters.
Discussion
The silencing or downregulation of tumor suppressor genes can happen both via genetic and epigenetic alterations. Most common epigenetic alteration such as aberrant promoter hypermethylation of TSG such as RASSF1a, APC1 and GSTP1 are reported in ovarian cancers.
RASSF1a is a tumor suppressor gene that is known to play a role in regulating several cellular mechanisms ranging from microtubule stabilization to apoptosis. We have reported here that 100% of samples in the benign and 93.75% low malignant potential tumors show down regulation of RASSF1a indicating that the silencing of RASSF1a is indeed an early event in the initiation and progression of ovarian cancer. In addition 94.1% of the malignant samples also show down regulation of RASSF1a. A similar down regulation of RASSF1a gene is also reported in cancers such as colon and prostate [27,28].
We have observed that the APC1 gene is downregulated in 77.77% and 56.25% of benign and low malignant potential tumors and the down regulation of APC gene was found to be statistically correlated with menopausal status. Several studies have reported the silencing of APC1 in prostate, colon, breast and ovarian cancer [28,30].
GSTP1 expression analysis revealed a 100% down regulation in benign, an 81.25% down regulation in LMPs and the silencing of GSTP1 correlated significantly with FIGO stage. The down regulation of GSTP1 is reported to contribute to the tumorigenesis of prostate, endometrial, hepatocellular, bladder and ovarian cancers [31-37].
Furthermore we have observed that 37 of the 76 samples analyzed showed a down regulation of all the three TSGs, suggesting that the silencing of RASSF1a, APC1 and GSTP1 are playing a crucial role in tumor initiation.
Development of molecular mechanism for the diagnosis of cancers is emerging as a promising method for several cancers. Assessment of mRNA expression of tumor suppressor genes can serve as a promising tool in diagnosis and management of ovarian cancer. Identifying and understanding the molecular mechanism contributing to the inactivation of TSG will aid in designing effective strategy for the treatment and management of the disease. Reversal of the silencing of tumor suppressor genes could further prove to be an effective mechanism in inhibiting cancer growth and metastasis.
Acknowledgements
The authors acknowledge Mr.Shanmugham V (Research Assistant, NIMHANS) for his guidance in statistical analysis of the data.
Declarations
Funding
SKS is the recipient of senior research fellowship from Council for Scientific and Industrial Research award no. 09/999/0002/2016-EMR-I. The research work was supported by Department of Biotechnology, India, Grant No BT/PR 13440/MED/30/267/2009 (2012).
Ethical Approval: The study was approved from the Institutional scientific and review board and medical ethics committee of Kidwai Memorial Institute of Oncology, Bangalore, India.
Conflict of Interest: The authors declare no conflict of interest.
References
- Lheureux S, Gourley C, Vergote I, Oza AM. Epithelial ovarian cancer. The Lancet 393 (2019): 1240-1253.
- Desai A, Xu J, Aysola K, et al. Epithelial ovarian cancer: An overview. World Journal of Translational Medicine 3 (2014): 1.
- Kumar S, Swamy SN, Premalatha CS, et al. Aberrant promoter Hypermethylation of RASSF1a and BRCA1 in circulating cell-free tumor DNA serves as a biomarker of ovarian carcinoma. Asian Pacific Journal of Cancer Prevention: APJCP 20 (2019): 3001.
- Swamy SN, Sandeep Kumar S, Devaraj VR, et al. Generation and Characterization of A Carboplatin Resistant Ovarian Cancer Model. Journal of Cancer Science and Clinical Therapeutics 4 (2020): 144-153.
- Weinberg RA. Tumor suppressor genes. Science 254 (1991): 1138-1146.
- Hinds PW, Weinberg RA. Tumor suppressor genes. Current Opinion in Genetics & Development 4 (1994): 135-141.
- Zhao M, Kim P, Mitra R, et al. TSGene 2.0: an updated literature-based knowledgebase for tumor suppressor genes. Nucleic Acids Research 44 (2016): D1023-D1031.
- Fiolka R, Zubor P, Janusicova V, et al. Promoter hypermethylation of the tumor-suppressor genes RASSF1A, GSTP1 and CDH1 in endometrial cancer. Oncology Reports 30 (2013): 2878-2886.
- Donninger H, Vos MD, Clark GJ. The RASSF1A tumor suppressor. Journal of Cell Science 120 (2007): 3163-3172.
- Chen H, Suzuki M, Nakamura Y, et al. Aberrant methylation of RASGRF2 and RASSF1A in human non-small cell lung cancer. Oncology Reports 15 (2006): 1281-1285.
- Honorio S, Agathanggelou A, Schuermann M, et al. Detection of RASSF1A aberrant promoter hypermethylation in sputum from chronic smokers and ductal carcinoma in situ from breast cancer patients. Oncogene 22 (2003): 147-150.
- Yazici H, Terry MB, Cho YH, et al. Aberrant methylation of RASSF1A in plasma DNA before breast cancer diagnosis in the Breast Cancer Family Registry. Cancer Epidemiology and Prevention Biomarkers 18 (2009): 2723-2725.
- Yoon JH, Dammann R, Pfeifer GP. Hypermethylation of the CpG island of the RASSF1A gene in ovarian and renal cell carcinomas. International Journal of Cancer 94 (2001): 212-217.
- Kawamoto K, Okino ST, Place RF, et al. Epigenetic modifications of RASSF1A gene through chromatin remodeling in prostate cancer. Clinical Cancer Research 13 (2007): 2541-2548.
- De Caceres II, Battagli C, Esteller M, et al. Tumor cell-specific BRCA1 and RASSF1A hypermethylation in serum, plasma, and peritoneal fluid from ovarian cancer patients. Cancer Research 64 (2004): 6476-6481.
- Giannopoulou L, Chebouti I, Pavlakis K, et al. RASSF1A promoter methylation in high-grade serous ovarian cancer: A direct comparison study in primary tumors, adjacent morphologically tumor cell-free tissues and paired circulating tumor DNA. Oncotarget 8 (2017): 21429.
- Karbova E, Davidson B, Metodiev K, et al. Adenomatous polyposis coli (APC) protein expression in primary and metastatic serous ovarian carcinoma. International Journal of Surgical Pathology 10 (2002): 175-180.
- Pellman D. A CINtillating new job for the APC tumor suppressor. Science 291 (2001): 2555-2556.
- van Es JH, Giles RH, Clevers HC. The many faces of the tumor suppressor gene APC. Experimental Cell Research 264 (2001): 126-134.
- Baeg GH, Matsumine A, Kuroda T, et al. The tumour suppressor gene product APC blocks cell cycle progression from G0/G1 to S phase. The EMBO Journal 14 (1995): 5618-5625.
- Morin PJ, Vogelstein B, Kinzler KW. Apoptosis and APC in colorectal tumorigenesis. Proceedings of the National Academy of Sciences 93 (1996): 7950-7954.
- Zhang L, Shay JW. Multiple roles of APC and its therapeutic implications in colorectal cancer. JNCI: Journal of the National Cancer Institute 109 (2017).
- Narayan S, Roy D. Role of APC and DNA mismatch repair genes in the development of colorectal cancers. Molecular Cancer 2 (2003): 41.
- Al-Shabanah OA, Hafez MM, Hassan ZK, et al. Methylation of SFRPs and APC genes in ovarian cancer infected with high risk human papillomavirus. Asian Pac J Cancer Prev 15 (2014): 25.
- Makarla PB, Saboorian MH, Ashfaq R, et al. Promoter hypermethylation profile of ovarian epithelial neoplasms. Clinical Cancer Research 11 (2005): 5365-5369.
- Bhagat R, Chadaga S, Premalata CS, et al. Aberrant promoter methylation of the RASSF1A and APC genes in epithelial ovarian carcinoma development. Cellular Oncology 35 (2012): 473-479.
- Zheng W, Zhao L, Wang G, et al. Promoter methylation and expression of RASSF1A genes as predictors of disease progression in colorectal cancer. International Journal of Clinical and Experimental Medicine 9 (2016): 2027-2036.
- Kaminska K, Bialkowska A, Kowalewski J, et al. Differential gene methylation patterns in cancerous and non-cancerous cells. Oncology Reports 42 (2019): 43-54.
- Makarla PB, Saboorian MH, Ashfaq R, et al. Promoter hypermethylation profile of ovarian epithelial neoplasms. Clinical Cancer Research 11 (2005): 5365-5369.
- Debouki-Joudi S, Trifa F, Khabir A, et al. CpG methylation of APC promoter 1A in sporadic and familial breast cancer patients. Cancer Biomarkers 18 (2017): 133-141.
- Lin X, Tascilar M, Lee WH, et al. GSTP1 CpG island hypermethylation is responsible for the absence of GSTP1 expression in human prostate cancer cells. The American Journal of Pathology 159 (2001): 1815-1826.
- Martignano F, Gurioli G, Salvi S, et al. GSTP1 methylation and protein expression in prostate cancer: diagnostic implications. Disease Markers 2016 (2016).
- Mian OY, Khattab MH, Hedayati M, et al. GSTP1 Loss results in accumulation of oxidative DNA base damage and promotes prostate cancer cell survival following exposure to protracted oxidative stress. The Prostate 76 (2016): 199-206.
- Chan QK, Khoo US, Chan KY, et al. Promoter methylation and differential expression of π-class glutathione S-transferase in endometrial carcinoma. The Journal of Molecular Diagnostics 7 (2005): 8-16.
- Li QF, Li QY, Gao AR, Shi QF. Correlation between promoter methylation in the GSTP1 gene and hepatocellular carcinoma development: a meta-analysis. Genet Mol Res 14 (2015): 6762-6772.
- Hadami K, Dakka N, Bensaid M, et al. Evaluation of glutathione S?transferase pi 1 expression and gene promoter methylation in Moroccan patients with urothelial bladder cancer. Molecular Genetics & Genomic Medicine 6 (2018): 819-827.
- Shilpa V, Bhagat R, Premalata CS, et al. GSTP1 expression and promoter methylation in epithelial ovarian carcinoma. Clinical Cancer Investigation Journal 3 (2014): 487.